February 19, 2019 – Machine learning, which provides the ability to learn a task from data (without the need of being programmed explicitly), is a key component of any Pathology AI (Artificial Intelligence) system. There are many different approaches in machine learning, reaching from simple decision trees to complex deep learning, each with its advantages and disadvantages.
Deep learning, which allows to learn highly complex visual features, has created a hype about Artificial Intelligence (AI) and Healthcare AI, as is was able to solve complex computer vision problems that we believed out-of-reach just a few years ago. As pathology is a visual task it is understandable that academia and “pure” technology companies are now working heavily on deep learning approaches for pathology. The key problem for any Pathology AI system are the variations between different patient types. In a disease state, no two patient samples look identical. To distinguish between different cell types, which any machine learning system has to accomplish somehow (even if it is hidden in some obscure features in a deep learning network), we notice that the same cell type has different characteristics in different patients, which are often contradictory.
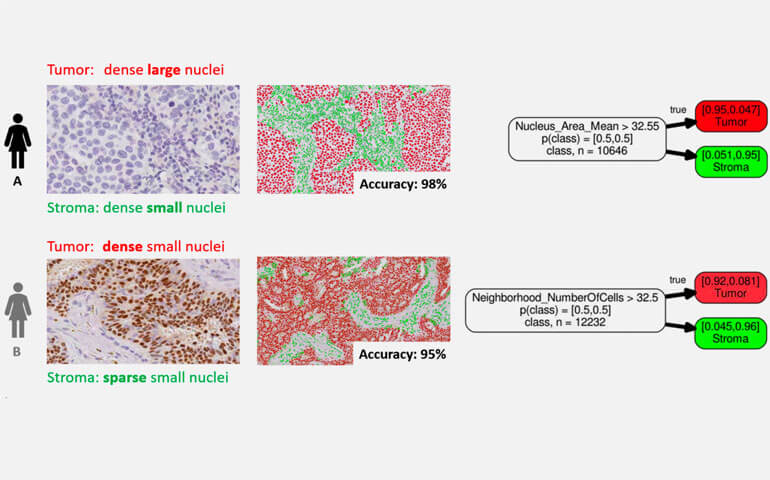
The example above shows two patients that are representative for two different patient types. Patient A has tumor cells that can be described as dense large nuclei and stroma cells that can be described as dense small nuclei. Patient B has tumor cells that can be described as dense small nuclei and stroma cells that can be described as sparse small nuclei. This illustrates nicely what we mean by contradictory characteristics. The stroma cells in patient A have the same characteristics as the tumor cells in patient B: dense small nuclei!
Let’s assume that our machine learning system consists of a single classifier that is pre-trained using a training set of histology slide images.
If we were to create a machine learning system that would be trained only on patients that have the same cell characteristics as patient A, the system would fail when it encounters a new patient type, like patient B. This illustrates the bias of machine learning systems that originates from the data used for training. We would need a lot of training data to make sure that the machine learning system would be able to learn the characteristics of all patient types properly. Getting a machine learning system from 90% performance to 95% or even 99% performance becomes exponentially harder as remaining exceptional cases are typically hard to come by.
Now if we were to create a machine learning system that would be trained on patients belonging to different patient types, like patient A and patient B, the system would have to learn somewhat “contradictory” data. We would need to use a complex machine learning approach that would be able to learn highly complex visual features with different contexts.
Obviously, deep learning would be right tool for that job.
Unfortunately, deep learning provides no transparency into the decision process, which eventually will have to face some legal and regulatory hurdles, as pathology is used to make medical decisions which put human lives at risk.
If the variations between different patient types is the key problem, let’s take that out of the equation.
Let’s go with a machine learning system which uses multiple “patient type”-specific classifiers that pathologists can train “on the fly”. See our article “Pathologist vs Artificial Intelligence (AI): Competition or collaboration” for a discussion on why you would like to have a pathologist as part of your machine learning system. The training ”on the fly” described here is how pathologists can best provide their expertise in a Pathology AI system.
In pathology there is no critical need to use machine learning to learn the visual features in histology slide. We are not looking at arbitrary objects in an uncontrolled environment, we are looking for cells that have a certain size and that have 3 cell compartments, the nucleus, the cytoplasm and the membrane and that can only be stained by a small number of different stains that have distinct colors. Traditional image analysis can do a good job detecting cells and measuring a wealth of biology motivated features that provide all the information there is in a histology slide.
Let’s go with a machine learning system which is based on cell data, not pixel data.
That kind of a machine learning system requires no training data, yields excellent performance and provides transparency into the decision process!
Here is how it works.
1) When we encounter the first patient, like patient A, or a patient that belongs to a new patient type (identified in 2), like patient B, we create a new “patient type”-specific classifier. A pathologist, using his expertise, identifies a few example regions for the different cell types (e.g. tumor and stroma) and trains a new classifier “on the fly” that then is used to classify all the cells on the whole slide. Proper controls are implemented by having the pathologist verify the proper classification of the cells. New example regions are added and the classifier is retrained until the pathologist is satisfied with the cell classification.
2) With any new patient, we first select the best classifier from all existing “patient type”-specific classifiers that then is used to classify all the cells on the whole slide. A very simple and robust method that nicely illustrate the selection of the best classifier is to have a pathologist just identify an example region for one (or more) cell type(s) and select the classifier that provides the best classification performance on those regions. When the pathologist now verifies the proper classification of the cells, he may decide that the classification is not good enough, which means that the new patient belongs to a new patient type and a new “patient-type” specific classifier needs to be created (go to 1).
The example above illustrates nicely that if we were to create different classifiers for different patient types, represented by patient A and patient B, that a simple decision tree using just a single feature with a single threshold would provide excellent performance and easy interpretable decisions. The results obtained by machine learning match nicely what we have seen by eye, the separation between tumor and stroma cells in patient A is based on nuclei size and in patient B on density of cells.
Limiting the machine learning to a specific patient type and using cell data simplifies the machine learning problem considerably, providing excellent performance with simple machine learning approaches like decision trees, which consist of easy to understand hierarchical flowcharts, and only requiring data from very few regions for training. The training is ultra-fast and can be done “on the fly” in an interactive and iterative workflow. A decision tree based on biology motivated features provides easy interpretable data to biologists and pathologists and a meaningful grouping of patients by patient type.
See our article “How Pathology AI works for Immuno-Oncology, a short demo” for a YouTube video on how that kind of a machine learning system works.
Holger Lange, CTO
Flagship Biosciences, Inc.